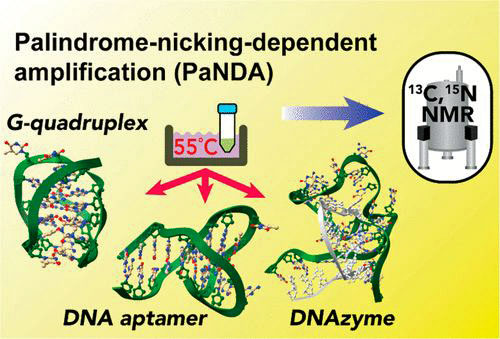
Single-stranded DNA (ssDNA) oligonucleotides are widely used in biological research, therapeutics, biotechnology, and nanomachines. Large-scale enzymatic production of ssDNA oligonucleotides forming noncanonical structures has been difficult.
In this webinar, Dr. Junji Iwahara presents a simple and robust method called “palindrome-nicking-dependent amplification” (PaNDA) for enzymatic production of a large amount of ssDNA oligonucleotides.
It utilizes a strand-displacing DNA polymerase and a nicking enzyme together with input DNA and deoxynucleotide triphosphates at 55 °C.
The 13C/15N-labeling by PaNDA greatly facilitates the characterization of noncanonical DNA by nuclear magnetic resonance (NMR) spectroscopy. For example, the behavior of therapeutic DNA aptamers in human serum can be investigated.
Reference: https://doi.org/10.1021/jacs.3c11219, J Am Chem Soc 2024, 146, in press
Watch the Webinar from January 31st 2024 :

Speaker: Dr. Junji Iwahara
Professor
University of Texas Medical Branch
Through his entire career, Prof. Junji Iwahara has studied various aspects of protein-DNA interactions using NMR spectroscopy and other biophysical methods.
More information about the speaker, Prof. Junji Iwahara
Visit the Iwaraha Lab Website
Education & Training
- 1989 – 1993 University of Tokyo (Japan): BS
- 1993 – 1998 University of Tokyo (Japan): PhD (Supervisor: Shigeyuki Yokoyama)
- 1998 – 2002 University of California, Los Angeles: Postdoctoral Fellow (Supervisor: Robert T. Clubb)
- 2002 – 2007 National Institutes of Health: Research Fellow(Supervisor: G. Marius Clore)
Faculty Positions
- 2017 – Present Dept. of Biochemistry & Molecular Biology, UTMB: Professor
- 2012 – 2017 Dept. of Biochemistry & Molecular Biology, UTMB: Associate Professor
- 2007 – 2012 Dept. of Biochemistry & Molecular Biology, UTMB: Assistant Professor
ORCiD: Junji Iwahara
Professor Iwahara’s current research interests:
Biomolecular Electrostatics
Professor Iwahara utilizes various Nuclear Magnetic Resonance (NMR) methods in combination with Stable Isotope Labeling approaches to quantitatively measure various biomolecular electrostatics even in the absence of full structural information.
Specifically, Professor Iwaara utilizes NMR spectroscopy for such characterizations of the positively or negatively charged groups of proteins and nucleic acids. 13C, 15N, and 31P nuclei that are useful for such studies: namely, 13C nuclei of aspartate/glutamate carboxylates, 15N nuclei arginine guanidinium cations, 15N nuclei of lysine amino cations; and 31P nuclei of phosphate anions of DNA, RNA, and phosphorylated protein side chains.
Selected publications
- Iwahara, J., Pettitt, B.M., Yu, B. (2023) Direct measurements of biomolecular electrostatics through experiments. Curr Opin Struct Biol 82, 102680.
- Yu, B., Pletka, C.C., Iwahara, J. (2022) Protein electrostatics investigated through paramagnetic NMR for nonpolar groups. J Phys Chem B 125, 2196-202.
Protein-DNA Dynamics
Proteins and DNA molecules are dynamic in many aspects. Using nuclear magnetic resonance (NMR) spectroscopy as our primary tool for research, we study various dynamics of proteins and DNA. Using these methodologies, we study how proteins scan DNA.
Selected publications
- Wang, X., Bigman, L., Greenblatt, H.M., Yu, B., Levy, Y., Iwahara, J. (2023) Negatively charged, intrinsically disordered regions can accelerate target search by DNA-binding proteins. Nucleic Acids Res 51, 4701-12
- Yu, B., Pletka, C.C., Iwahara, J. (2021) Quantifying and visualizing weak interactions between anions and proteins. Proc Natl Acad Sci U S A 118 (2), e2015879118.
HMGB1 Biochemistry
HMGB1 is a very interesting DNA-binding protein that plays important roles in the cell nuclei as well as in extracellular space. HMGB1 typically exists in the cell nuclei and serves as a DNA chaperone and assists various other DNA-binding proteins. HMGB1 contains two DNA-binding domains and intrinsically disordered regions (IDRs).
HMGB1 recognizes atypical DNA such as Holliday junction, bulged DNA, cisplatin-modified DNA, and G-quadruplex. HMGB1 is released to extracellular space not only passively from dying cells, but also actively from platelets and some healthy cells. Extracellular HMGB1 serves as a danger signal and plays several important roles in innate immunity involving extracellular DNA related to infection or tissue damage. HMGB1 is involved in many inflammatory diseases and has been regarded as a potential therapeutic target.
Using NMR spectroscopy, fluorescence spectroscopy, and other techniques, we are characterizing HMGB1 and trying to find effective ways for HMGB1 inhibition.
- Wang, X., Bigman, L., Greenblatt, H.M., Yu, B., Levy, Y., Iwahara, J. (2023) Negatively charged, intrinsically disordered regions can accelerate target search by DNA-binding proteins. Nucleic Acids Res 51, 4701-12.
- Wang, X., Mayorga-Flores, M., Bien, K., Bailey, A.O., Iwahara, J. (2022) DNA-mediated proteolysis by neutrophil elastase enhances binding activities of the HMGB1 protein. J Biol Chem 298, 102577.